Introduction
Human cells become cancerous when they undergo genetic modifications that make them acquire growth and multiplication advantages. Cancerous cells do not carry out normal physiological roles in human beings. The processes that culminate in genetic changes occur in an orderly manner that involves the accumulation of inherited changes that alter the functions of genes. Two classes of genes have been implicated in the development of cancers in human beings (Huang, Nayak, Jankowitz, Davidson & Oesterreich 2011). First, tumour suppressor genes prevent cells from growing and surviving. Second, oncogenes facilitate the processes of cell growth and survival. Research has shown that different mechanisms are involved in converting cells from normal conditions to cancerous states (Grønbæk, Hother & Jones 2007). Grønbæk and colleagues (2007) assert that the mechanisms of cell alterations ultimately modify the proteins that encode nucleotides, alter the number of genes and increase the rate of gene transcription. Signalling pathways in cells are important in maintaining normal cell functions. However, epigenetic and genetic changes have been shown to cause abnormal cell activities by interfering with the normal pathways involved in cell functions. The number of oncogenes could be increased by gaining extra number of chromosomes, amplifying genes, activating point mutations and shifting the building blocks of DNA. Also, tumour suppressor genes could be deactivated by deleting gross regions in chromosomes, deleting small gene portions and deleting specific nucleotides within a DNA sequence.
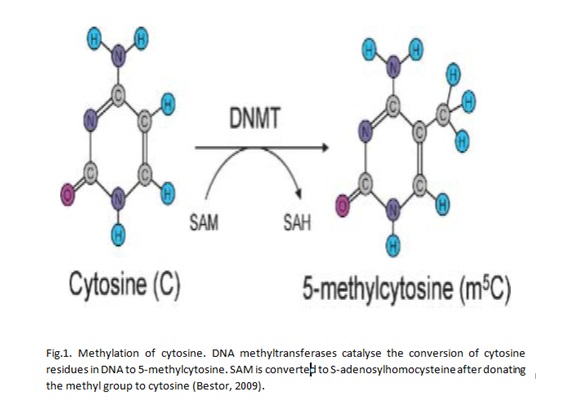
Epigenetics is the study of how cell activities are inherited through DNA and RNA patterns that are not influenced by nucleotide sequences. Methylation is an example of epigenetic change that has been widely studied. DNA methylation occurs after DNA replication and it is characterised by the addition of methyl groups to DNA molecules at the sites with cytosine residues. Research has shown that cytosine residues are vey close to CpG islands that are marked by a high concentration of guanidine residues (Huang et al 2011). Thus, methylation of DNA occurs in regions with a high concentration of guanidine residues. Bird (2002) asserts that CpG islands are found in about two thirds of gene promoters. Genes with DNA that is hypermethylated are not expressed. On the other hand, hypomethylation is correlated with increased gene activities. Thus, it could be expected that cancer cells have DNA molecules that are hypomethylated. Specifically, oncogenes are hypomethylated while tumour suppressor genes are characterised by high levels of hypermethylation. Thus, it could be asserted that DNA methylation has implications for the development and progression of cancer.
DNA methylation
Research has identified three enzymes that are associated with DNA methylation in human beings. First, DNA methyltransferase 1 (DNMT1) is involved in maintaining methylation patterns in cells. Second, DNA methyltransferase 3A (DNMT3A) is concerned with regulating de novo methylation processes in cells. It has also been suggested that DNA methyltransferase 3B (DNMT3B) has the same functions as that of DNMT3A (Chen et al 2007; Chedin 2010). The three enzymes are expressed at different rates, but their modes of expression do not always correlate with hypomethylation and hypermethylation. The variations are caused by the action of miRNAs that regulate the molecular events of DNMT (Blair and Yan 2012).
There are different mechanisms for DNA methylation. One of the best explained mechanisms is through the involvement of TET1-3 proteins, which belong to hydroxylases (Tahiliani et al 2009). The molecules catalyse the conversion of 5mc, 5HMC, 5fC and 5caC (Ito, D’Alessio, Taranova, Hong, Sowers & Zhang 2010; He et al 2011; Pfaffeneder et al 2011).
Point mutations due to DNA methylation
Grønbæk and colleagues (2007) show that point mutations are promoted in regions characterised by 5mc residues through many ways. A cytosine that could be methylated could encounter deamination to form a thymine. If the changes are not corrected by independent molecular events in cells, then they result in disease due to point mutations in regions coding for genes. In most cases, genes that are affected are those that are concerned with the regulation of cell growth and survival. More than 30% of human diseases that are associated with point mutations have been shown to have alterations at the CpG dinucleotides.
Chromatin and histone modification
Chromatin is the physiological template for the human genome. The basic unit of chromatin is the nucleosome that is characterised by about 200bp of DNA. The base pairs are organised in small basic units of proteins that exist as octamers. A collection of octamers is known as histone. Thus, histone is the major molecular material that is involved in organising and maintaining the structural integrity of DNA and genes. The human DNA has been found to lie on the surface while histone materials are found in the inner parts of histone molecules. Nucleosomes are part of euchromatin and heterochromatin. Also, euchromatin and heterochromatin are found in mitotic chromosomes. However, heterochromatin and euchromatin of the human genome differ significantly in terms of structure and function. Heterochromatin is packed tightly to prevent transcription factors from accessing histone. On the other hand, euchromatin is not tightly packed. Thus, factors involved in transcription could easily access major regions of DNA that are involved in the initiation of transcription of DNA (Richards & Elgin 2002).
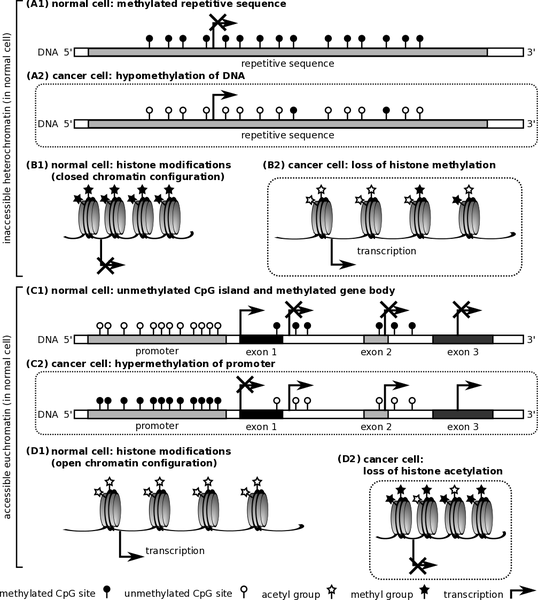
Research has also shown that cancer could be caused by epigenetic modifications that alter the normal structure of histone. DNA molecules in every cell nucleus wraps around histone materials, which have an extensive altered N-terminal tails that could also be flexible. The rate at which transcription of DNA occurs greatly depends on specific alterations that could either tighten or loosen molecules of DNA from histone (Blair and Yan 2012).
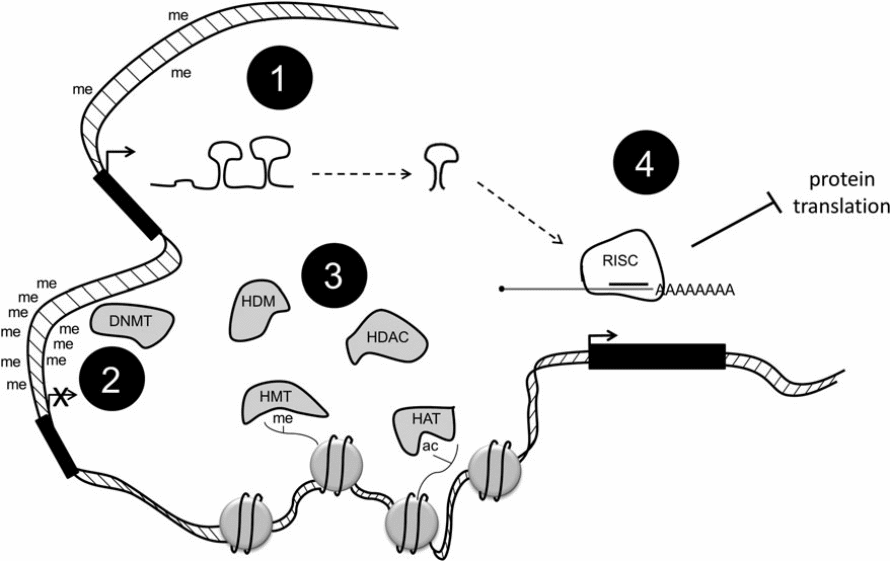
Some alterations of histone correlate with active genes such as trimethylation in histone H3. Some changes could also be connected with genes that could be repressed such as H3K9me3. Acetylation of histone occurs when histone acetyltransferases (HATs) catalyse the addition of acetyl groups to lysine residues. The remove of such residue is catalysed by histone deacetylase (HDACs). Methyl groups are removed through the action of histone demethylases (HDMs). Another important modification of histone is phosphorylation, which ensures that essential residues in histone are phosphorylated. The roles of the enzymes that catalyse the activities of phosphorylation, methylation, and acetylation are not clearly understood (Blair & Yan 2012).
Breast cancer
Research shows that BRAC1 and BRAC2 genes could be responsible for coding proteins that make breast cells grow and multiply abnormally. Disease progression and degrees of severity vary among the different types of breast cancer (Stecklein, Jensen & Pal 2012). If a patient lacks the three receptors that signify the presence of breast cancer, then such a patient is said to be triple negative. It has been shown that a patient presenting with cancer characterised by this state (triple negative) could be difficult to treat. In such patients, there are no readily available hormonal therapy approaches that could be used to treat cancer. Patients who present with cancers characterised by estrogen receptor (ER) tumours are treated using hormonal therapy. Clinical data also show that trastuzumab (Herceptin®) has been used to treat patients with human epidermal growth receptor (HER) positive tumours (Blair & Yan 2012). Epidermal growth factor has been correlated with cancers with high rates of metastasis and invasiveness. Such tumours are quite difficult to treat, especially when they are treated at advanced stages. At such advanced stages, the tumours could have spread to other body organs where they cause physiological damage. However, the effect of EGFR on the development and progression of breast cancers has been reduced through the use of tyrosine kinase inhibitors. An example of such inhibitors is gentinib. The pharmacological product has also been shown to target cells with over-expression of HER2.
Many forms of breast cancer are hypomethylated (Soares, Pinto, Cunha, Andre, Barão, Sousa & Cravo 1999). The observation is similar to findings from other studies (Hinshelwood & Clark 2008). Hypermethylation in noncancerous cells does not occur in the same regions like those in cancer cells (Hinshelwood & Clark 2008). This is a major observation that differentiates molecular events between caner and healthy cells. In order for cancer to progress, tumour suppressor genes need to be suppressed. If they are suppressed, then they lose the ability to prevent the growth of tumours associated with the progression of cancer. DNA hypermethylation results in unstable genes that could lead to cancer. In fact, a significant number of unstable genes results in various forms of cancer because they code for abnormal proteins that facilitate cells to grow abnormally.
DNMTs could be involved in gene-based methylation that is common in breast cancer (Girault, Tozlu, Lidereau & Bièche 2003). However, a weak correlation between DNMTs and breast cancer has been demonstrated. Methylation of the promoter of ER could result in the down-regulation of ER. Breast cancers with over-expressed ER and PR are treated through the use of drugs that imitate the receptors (Blair & Yan 2012). Scientists have concentrated on deciphering the roles of HMTs and HDMs in the development and progression of breast cancer. Such efforts have shown that there are elevated levels of H3K27 methyltransferases, which enhance the activities of zeste homologue 2 (EZH2). Cancer states that are characterised by elevated levels of H3K27 methyltransferases have fast rates of metastasis. Such cancers are difficult to treat and manage. Breast cancer is also associated with over-expression of lysine-specific demethylases that are involved in the removal of methyl groups from lysine residues. Research has demonstrated that there is over-expression of H3K4 demethylase in ER negative tumours. In fact, this is being used as a diagnostic marker for breast cancers that have high chances of metastasising.
The discovery of the genes and molecules involved in the development and progression of breast cancers has been very sequential. The orderly manner in which scientists have been discovering breast cancer-specific genes and molecules has greatly impacted the management of the disease. For example, hormonal therapy has been adopted in the treatment of breast cancers that are characterised by ER. In the future, further research could result in the identification of more genes and molecules with regard to breast cancer. Therefore, more treatment approaches would be designed in the future.
Aim and objective
DNMT1 is essential in maintaining DNA methylation pathways. If the enzyme is over-expressed, then hypermethylation occurs. This is the implication for many forms of breast cancer (Girault et al 2003). The proposed study aims at assessing the effect of demethylation on the growth of normal human cells and development and progression of breast cancer cells. The study will use the wound healing method that is often used in the study of the biology involved in the migration of cells (Francisco et al 2003).
Hypothesis
The study hypothesises that a significant change (arrest of cell migration) will be observed when breast cancer cell lines will be demethylated using a demethylating enzyme.
Materials and methods
Assays will be conducted in a biotechnology laboratory that will have all the required facilities.
Cell lines
The MCF-10A and HMT-3522 will be the normal epithelial cells and the T47D and ZR-75-1 will be breast cancer cell lines that will be utilised in the study.
Protocol
A standard protocol will be used to maintain the cell lines in vitro. All cell lines will be allowed to grow in independent dishes for a period of 48 hours. The dishes will be supplied with RPMI-1640 that will contain essential growth materials such as L-glutamate, penicillin, streptomycin and 50% carbon dioxide. A temperature of 370C will be maintained. DNMT1 (methylase) will be added to the normal epithelial cells while 5-methylcytosine (demethylase) will be added to breast cancer cell lines. The cells will then be monitored for a period of 24 hours. Controls will not have either a demethylating or methylating enzyme added. A wound will be stimulated by scratching the cell structures in the plates at an angle of 30 degrees. A sterile needle will be used. After 24 hours, cells will be washed using HBSS. Images will be taken every six hours to observe cell migration patterns. The metric relative wound density method will be utilised to determine the rate of cell migration. Afterwards, an automated Incucyte FLR machine will be used to achieve live cell imaging.
Analysis
The images that will be obtained from the cell migration (wound healing) assay will be studied through the use of MVTec software. In order to obtain a graph, the relative wound density assay figures will be plotted against time take to achieve specific distances of cell movements.
Expenditure of main reagents
Figure 4. Table showing the reagents to be used in the study and their prices.
Health and safety
Health and safety are important when conducting research activities in scientific laboratories. Precautions will be taken so that the various chemicals in the laboratory cannot cause harm to the personnel working in the laboratory. Standard health and safety procedures have been developed to help tom protect personnel in laboratories from any harm. Also, the standard procedures aim at protecting the environment from the negative impact of laboratory chemicals. Health and safety procedures will be used at all times while conducting the proposed study. Laboratory coats and gloves will be used. Contaminated materials will be decontaminated using standard procedures.
References
Bird, A, 2002, ‘DNA methylation patterns and epigenetic memory’, Genes & development, vol. 16, no. 1, pp. 6-21.
Blair, LP, & Yan, Q, 2012, ‘Epigenetic mechanisms in commonly occurring cancers’, DNA and cell biology, vol. 31, no. 1, pp. 49-61.
Chedin, F, 2010, ‘The DNMT3 family of mammalian de novo DNA methyltransferases’, Progress in molecular biology and translational science, vol. 101, no.1, pp. 255-285.
Chen, T, Hevi, S, Gay, F, Tsujimoto, N, He, T, Zhang, B.,… & Li, E, 2007, ‘Complete inactivation of DNMT1 leads to mitotic catastrophe in human cancer cells’, Nature genetics, vol. 39, no. 3, pp. 391-396.
Francisco, J. A., Cerveny, C. G., Meyer, D. L., Mixan, B. J., Klussman, K., Chace, D. F.,… & Wahl, A. F. (2003). cAC10-vcMMAE, an anti-CD30–monomethyl auristatin E conjugate with potent and selective antitumor activity. Blood, 102(4), 1458-1465.
Girault, I, Tozlu, S, Lidereau, R, & Bièche, I, 2003, ‘Expression analysis of DNA methyltransferases 1, 3A, and 3B in sporadic breast carcinomas’, Clinical Cancer Research, vol. 9, no. 12, pp. 4415-4422.
Grønbæk, K, Hother, C, & Jones, PA, 2007, ‘Epigenetic changes in cancer. Apmis, vol. 115, no. 10, pp. 1039-1059.
He, YF, Li, BZ, Li, Z, Liu, P, Wang, Y, Tang, Q,… & Xu, GL, 2011, ‘Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA’, Science, vol. 333, 6047, pp. 1303-1307.
Hinshelwood, R. A., & Clark, S. J. (2008). Breast cancer epigenetics: normal human mammary epithelial cells as a model system. Journal of molecular medicine, 86(12), 1315-1328.
Huang, Y, Nayak, S, Jankowitz, R, Davidson, NE, & Oesterreich, S, 2011, ‘Epigenetics in breast cancer: what’s new?’, Breast Cancer Research, vol. 13, no. 6, pp. 1-11.
Ito, S, D’Alessio, AC, Taranova, OV, Hong, K, Sowers, LC, & Zhang, Y, 2010, ‘Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification’, Nature, vol. 466, no. 7310, pp. 1129-1133.
Pfaffeneder, T, Hackner, B., Truß, M, Münzel, M, Müller, M, Deiml, CA,… & Carell, T, 2011, ‘The discovery of 5‐formylcytosine in embryonic stem cell DNA’, Angewandte Chemie, vol. 123, no. 31, pp. 7146-7150.
Richards, EJ, & Elgin, SC, 2002, ‘Epigenetic codes for heterochromatin formation and silencing: rounding up the usual suspects’, Cell, vol. 108, no. 4, pp. 489-500.
Soares, J, Pinto, AE., Cunha, CV, Andre, S, Barão, I, Sousa, JM, & Cravo, M, 1999, ‘Global DNA hypomethylation in breast carcinoma’, Cancer, vol. 85, no. 1, pp. 112-118.
Stecklein, SR., Jensen, RA, & Pal, A, 2012, ‘Genetic and epigenetic signatures of breast cancer subtypes’, Front Biosci (Elite Ed), vol. 4, no. 1, pp. 934-949.
Tahiliani, M, Koh, KP, Shen, Y, Pastor, WA, Bandukwala, H, Brudno, Y,… & Rao, A, 2009, ‘Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1’, Science, vol. 324, no. 5929, pp. 930-935.